Utility Functions¶
A group of useful functions
-
modred.util.Hankel(first_col, last_row=None)[source]¶ Construct a Hankel array, whose skew diagonals are constant.
- Args:
first_col: 1D array corresponding to first column of Hankel array.- Kwargs:
last_row: 1D array corresponding to the last row of Hankel array. First element will be ignored. Default is an array of zeros of the same size asfirst_col.- Returns:
- Hankel: 2D array with dimensions
[len(first_col), len(last_row)].
-
modred.util.Hankel_chunks(first_col_chunks, last_row_chunks=None)[source]¶ Construct a Hankel array using chunks, whose elements have Hankel structure at the chunk level (constant along skew diagonals), rather than at the element level.
- Args:
first_col_chunks: List of 2D arrays corresponding to the first column of Hankel array chunks.- Kwargs:
last_row_chunks: List of 2D arrays corresponding to the last row of Hankel array chunks. Default is a list of arrays of zeros.- Returns:
- Hankel: 2D array with dimension
[len(first_col) * first_col[0].shape[0], len(last_row) * last_row[0].shape[1]].
-
class
modred.util.InnerProductBlock(inner_product)[source]¶ Only used in tests. Takes inner product of all vectors.
-
modred.util.atleast_2d_col(array)[source]¶ Converts 1d arrays to 2d arrays, but always as column vectors
-
modred.util.atleast_2d_row(array)[source]¶ Converts 1d arrays to 2d arrays, but always as row vectors
-
modred.util.balanced_truncation(A, B, C, order=None, return_sing_vals=False)[source]¶ Balance and truncate discrete-time linear time-invariant (LTI) system defined by A, B, C arrays. It’s not very accurate due to numerical issues.
- Args:
A,B,C: LTI discrete-time arrays.- Kwargs:
order: Order (number of states) of truncated system. Default is to use the maximal possible value (can truncate system afterwards).- Returns:
A_balanced,B_balanced,C_balanced: LTI discrete-time arrays of balanced system.If
return_sing_valsis True, also returns:sing_vals: Hankel singular values.
Notes:
Dis unchanged by balanced truncation.- This function is not computationally efficient or accurate relative to
Matlab’s
balancmr.
-
modred.util.drss(num_states, num_inputs, num_outputs)[source]¶ Generates a discrete-time random state-space system.
- Args:
num_states: Number of states.num_inputs: Number of inputs.num_outputs: Number of outputs.- Returns:
A,B,C: State-space arrays of discrete-time system.
By construction, all eigenvalues are real and stable.
-
modred.util.eig_biorthog(array, scale_choice='left')[source]¶ Wrapper for
numpy.linalg.eigthat returns both left and right eigenvectors. Eigenvalues and eigenvectors are sorted and scaled so that the left and right eigenvector arrays are orthonormal.- Args:
array: Array to take eigendecomposition of.- Kwargs:
scale_choice: Determines whether ‘left’ (default) or ‘right’ eigenvectors will be scaled to yield a biorthonormal set. The other eigenvectors will be left unscaled, leaving them with unit norms.- Returns:
evals: 1D array of eigenvalues.R_evecs: Array whose columns are right eigenvectors.L_evecs: Array whose columns are left eigenvectors.
-
modred.util.eigh(array, atol=1e-13, rtol=None, is_positive_definite=False)[source]¶ Wrapper for
numpy.linalg.eigh. Computes eigendecomposition of a Hermitian array.- Args:
array: Array to take eigendecomposition of.atol: Value below which eigenvalues (and corresponding eigenvectors) are truncated.rtol: Maximum relative difference between largest and smallest eigenvalues. Smaller ones are truncated.is_positive_definite: If true, array being decomposed will be assumed to be positive definite. Tolerance will be automatically adjusted (if necessary) so that only positive eigenvalues are returned.- Returns:
eigvals: 1D array of eigenvalues, sorted in descending order (of magnitude).eigvecs: Array whose columns are eigenvectors.
-
modred.util.get_file_list(directory, file_extension=None)[source]¶ Returns list of files in
directorywithfile_extension.
-
modred.util.impulse(A, B, C, num_time_steps=None)[source]¶ Generates impulse response outputs for a discrete-time system.
- Args:
A,B,C: State-space system arrays.- Kwargs:
num_time_steps: Number of time steps to simulate.- Returns:
outputs: Impulse response outputs, with indices corresponding to [time step, output, input].
No D array is included, but one can simply be prepended to the output if it is non-zero.
-
modred.util.load_array_text(file_name, delimiter=None, is_complex=False)[source]¶ Reads data saved in a text file, returns an array.
- Args:
file_name: Name of file from which to load data.- Kwargs:
delimiter: Delimiter in file. Default is same asnumpy.loadtxt.is_complex: Boolean describing whether the data to be loaded is complex valued.- Returns:
array: 2D array containing loaded data.
See
save_array_text()for the format used by this function.
-
modred.util.load_multiple_signals(signal_paths, delimiter=None)[source]¶ Loads multiple signal files from text files with columns [t channel1 channel2 …].
- Args:
signal_paths: List of filepaths to files containing signals.- Returns:
time_values: 1D array of time values.all_signals: Array of signals with indices [path, time, signal].
See
load_signals().
-
modred.util.load_signals(signal_path, delimiter=None)[source]¶ Loads signals from text files with columns [t signal1 signal2 …].
- Args:
signal_paths: List of filepaths to files containing signals.- Returns:
time_values: 1D array of time values.signals: Array of signals with dimensions [time, signal].
Convenience function. Example file has format:
0 0.1 0.2 1 0.2 0.46 2 0.2 1.6 3 0.6 0.1
-
modred.util.lsim(A, B, C, inputs, initial_condition=None)[source]¶ Simulates a discrete-time system with arbitrary inputs.


- Args:
A,B, andC: State-space system arrays.inputs: Array of inputs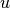 , with dimensions
, with dimensions
[num_time_steps, num_inputs].- Kwargs:
initial_condition: Initial condition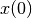 .
.- Returns:
outputs: Array of outputs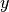 , with dimensions
, with dimensions
[num_time_steps, num_outputs].
Darray is assumed to be zero.
-
modred.util.make_iterable(arg)[source]¶ Checks if
argis iterable. If not, makes it a one-element list. Otherwise returnsarg.
-
modred.util.rss(num_states, num_inputs, num_outputs)[source]¶ Generates a continuous-time random state-space system.
- Args:
num_states: Number of states.num_inputs: Number of inputs.num_outputs: Number of outputs.- Returns:
A,B,C: State-space arrays of continuous-time system.
By construction, all eigenvalues are real and stable.
-
modred.util.save_array_text(array, file_name, delimiter=None)[source]¶ Saves a 1D or 2D array to a text file.
- Args:
array: 1D or 2D or array to save to file.file_name: Filepath to location where data is to be saved.- Kwargs:
delimiter: Delimiter in file. Default is same asnumpy.savetxt.
Format of saved files is:
2.3 3.1 2.1 ... 5.1 2.2 9.8 ... 7.6 3.1 5.5 ... 0.1 1.9 9.1 ... ...
Complex data is saved in the following format (as floats):
real[0,0] imag[0,0] real[0,1] imag[0,1] ... real[1,0] imag[1,0] real[1,1] imag[1,1] ... ...
Files can be read in Matlab with the provided functions or often with Matlab’s
load.
-
modred.util.smart_eq(arg1, arg2)[source]¶ Checks for equality, accounting for the fact that numpy’s
==doesn’t return a bool. In that case, returns True only if all elements are equal.
-
modred.util.sum_lists(list1, list2)[source]¶ Sums the elements of each list, returns a new list.
This function is used in MPI reduce commands, but could be used elsewhere too.
-
modred.util.svd(array, atol=1e-13, rtol=None)[source]¶ Wrapper for
numpy.linalg.svd, computes the singular value decomposition of an array.- Args:
array: Array to take singular value decomposition of.- Kwargs:
atol: Level below which singular values are truncated.rtol: Maximum relative difference between largest and smallest singular values. Smaller ones are truncated.- Returns:
U: Array whose columns are left singular vectors.S: 1D array of singular values.V: Array whose columns are right singular vectors.
Truncates
U,S, andVsuch that the singular values obey bothatolandrtol.